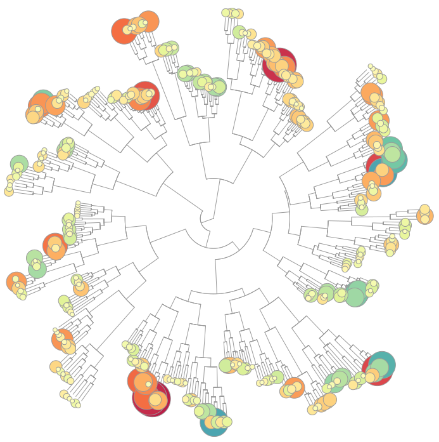
An R-Shiny-based implementation of weighted gene co-expression network analysis (WGCNA) obtained from the Primary Human Hepatocytes (PHH) TG-GATEs dataset.
<script> function initTeSSWidgets() { var query = ''; if (query.trim() != "") { TessWidget.Materials(document.getElementById('tess-widget-materials-list'), 'SimpleList', { opts: { enableSearch: false }, params: { pageSize: 5, q: query } }); document.getElementById('tess-widget-materials-header').innerHTML = "Documentation from ELIXIR TeSS" } } </script> <script async="" defer="" src="https://elixirtess.github.io/TeSS_widgets/components/js/tess-widget-standalone.js" onload="initTeSSWidgets()"></script>- External cloud: https://txg-mapr.eu/
- Login required: Yes
- Implementation status:
- TRL:
- Type: -
- Contact: Steven J. Kunnen [email protected]
- API Type:
- Categories: -
- Targeted users: -
- Relevant VHP4Safety Use case: -
- Provided by: Leiden University
- Citation: https://doi.org/10.1007/s00204-021-03141-w
- Version:
- License: Proprietary
- Source Code:
- Docker:
- Bio.tools: https://bio.tools/TXG-MAPr
- FAIRsharing:
- TeSS:
- RSD:
- Wikipedia: